Важным этапом в предиктивной медицине является этический вопрос о том, как оценивать абсолютные и относительные риски и как сообщать об этих рисках, чтобы наилучшим образом обслуживать каждого пациента – например, поощрять принятие изменений образа жизни или проводить скрининг заболеваний [1]. Кроме того, ассоциативные исследования выявили возможности перепрофилирования лекарств, позволяющие использовать препараты, уже одобренные по конкретным показаниям, для лечения альтернативных патогенных мишеней [2].
NGS как метод поиска причин репродуктивных потерь
Сегодня NGS биоматериала у пациентов с предполагаемым бесплодием позволяет провести общегеномные исследования ассоциаций, специфические для различных фенотипов бесплодия, таких, как предрасположенность к преждевременной недостаточности яичников, повышенному риску анеуплоидий, оценке незрелости яйцеклеток или недостаточности развития бластоцисты, что приведет к разработке настоящей репродуктивной точной медицины [3].
Бесплодие является одним из наиболее распространенных патологических состояний среди лиц в возрасте от 20 до 45 лет, поражающим 10–15% супружеских пар [4]. Приводятся оценки, что использование NGS до зачатия потенциально позволяет выявить наследуемые генетические заболевания, скрытые в геноме примерно у 4% пар, желающих забеременеть. Геномная информация в репродуктивном возрасте также будет полезна для прогнозирования поздних, поддающихся медикаментозному лечению состояний с сильным генетическим фоном, встречающимся примерно у 2–4% всех людей.
Существует несколько подходов применения технологии NGS для анализа репродуктивных потерь в семьях: а) экзомное (таргетное) секвенирование проводят обоим супругам с целью поиска мутаций, приводящих к задержке раннего эмбрионального и фетального развития (в гетерозиготном варианте); мутации в гомозиготе в этих же генах ищут в абортивном материале; б) проводят экзомное секвенирование абортивного материала с целью поиска рецессивных вариантов мутаций у выкидыша; найденные варианты могут дополнительно анализировать методом секвенирования по Сэнгеру у родителей или в другом абортивном материале; в) экзомное секвенирование с целью поиска гетерозиготного носительства в гене-кандидате проводят у одного родителя; в случае обнаружения методом Сэнгера ищут данную мутацию (в гомозиготе) в абортивном материале и в гетерозиготе у второго родителя; г) экзомное секвенирование с целью поиска маркера неразвивающейся беременности проводят только у матери [5].
Применение первого подхода демонстрирует исследование Filges I. et al. [6], в котором при тестировании трио (мать, отец и выкидыш) удалось подтвердить, что причиной неразвивающейся беременности стали мутации в гене KIF14, обнаруженные в компаунде в абортивном материале плода с микроцефалией, почечными и скелетными аномалиями. Данная находка была позже подтверждена при исследовании материала второго выкидыша, а роль продукта гена KIF14 в развитии мозга и почек была доказана в нокаутной модели рыбки Данио–Рерио [7]. В рамках второго подхода в качестве причины привычного невынашивания была идентифицирована компаудная мутация в гене ALOX15 [8]. Этот же подход применили и для установления причины рецидивирующих выкидышей мужского пола у плодов с водянкой и акинезией [9]. В качестве причины была идентифицирована мутация в гене FOXP3 на Х-хромосоме, связанная с аутоиммунным процессами. Мутация была подтверждена секвенированием по Сэнгеру у матери. Еще в одном исследовании причину выкидышей удалось установить, как мутацию в гене THSD1, продукт которого принимает участие в ангиогенезе [10]. В целом подобный подход позволил сконцентрировать внимание на более чем 286 генах-кандидатах ранней эмбриональной летальности [11]. Третий подход позволил идентифицировать еще несколько генов, мутации в которых имеют значение для ранней потери беременности, – GLE1, RYR1, ATRX. Более того, было продемонстрировано, что экзомное секвенирование позволяет детектировать не только однонуклеотидные замены – в одной семье причиной выкидыша было сочетание большой делеции в гене IFT122 и SNP [12]. В рамках четвертого подхода внимание исследователей сосредоточено на 234 генах кандидатах, связанных напрямую с процессами, проходящими при беременности – инвазией клеток трофобласта в материнскую ткань, коагуляцией, ремоделированием внеклеточного матрикса, миграцией, апоптозом, имунными процессами, активаций стероидных гормонов. В рамках данных исследований было выявлено два гена, в которых были обнаружены патогенные мутации, – FGA (коагуляция) и MMP10 (внеклеточный матрикс) [13].
Использование двух технологий – «низкопокрытого» генома и хромосомного микроматричного анализа (ХМА) позволило найти 275 потенциальных генов и 154 CNV, отвечающих за ранние репродуктивные потери [14]. Несмотря на то что трансцервикальная эмбриоскопия абортивного материала и трансвагинальное ультразвуковое исследование информируют нас об анатомических особенностях и деталях развития эмбриона, представляется, что использование кариотипирования в комбинации с NGS может существенно улучшить понимание причин ранних репродуктивных потерь [15].
Сложности применения NGS в репродукции
Расширение анализа секвенирования на дополнительные моногенные и полигенные признаки может позволить разработать экономически эффективные тесты до зачатия, способные идентифицировать основные генетические причины бесплодия, которые до сих пор определялись как «необъяснимые», что приведет к разработке действительно персонализированной системы геномной медицины в области репродуктивного здоровья.
Основные сложности в применении методов NGS на практике заключаются в следующем: данная технология выявляет огромное количество вариантов генов (более 20 тыс. на экзом), потенциально связанных с репродуктивными потерями, и задача врача-генетика, генетического консультанта и биоинформатика во многом сводится к аккуратному использованию биоинформатического протокола и имеющихся баз данных о патогенности и популяционных частотах встречаемости тех или иных вариантов. Общепринятым является подход, когда обращают внимание на патогенные варианты с частотой не более 1%. Интересно отметить, что бесплодие, по-видимому, не связано с повышенным выявлением патогенных вариантов в 59 вторичных находках списка генов ACMG [3]. В связи с чем, применение экзома выглядит более перспективным для генетического скрининга бесплодия и хронических моногенных заболеваний.
Репродуктивные потери и моногенные болезни
По оценкам, более 7000 менделевских или моногенных расстройств в совокупности влияют на 1–3% живорождений. С точки зрения здравоохранения, эти состояния оказывают гораздо большее влияние, поскольку они соответствуют гораздо большей доле общей заболеваемости и смертности (например, до 71% случаев госпитализации в педиатрические больницы) [16]. Подсчитано, что одно из 1300 известных на сегодняшний день рецессивных генетических заболеваний поражает минимум трех детей на каждые 1000. На самом деле, болезни с рецессивным наследованием могут быть относительно редкими, если рассматривать их по отдельности, но при совместном учете они становятся эпидемиологически значимыми [3].
Наиболее часто встречающиеся наследственные моногенные болезни с рецессивным типом наследования – муковисцидоз, фенилкетонурия и спинальная мышечная атрофия, хотя сегодня описано уже более 9000 наследственных заболеваний, которые так или иначе связаны с аномалиями в генах, и каждый год список увеличивается на 200–300 новых диагнозов.
Выделяют следующие группы моногенных заболеваний, связанные с репродуктивными потерями: моногенные заболевания матери (таблица), моногенные заболевания плода; другие генетические факторы (гены-предрасположенности, РНК).
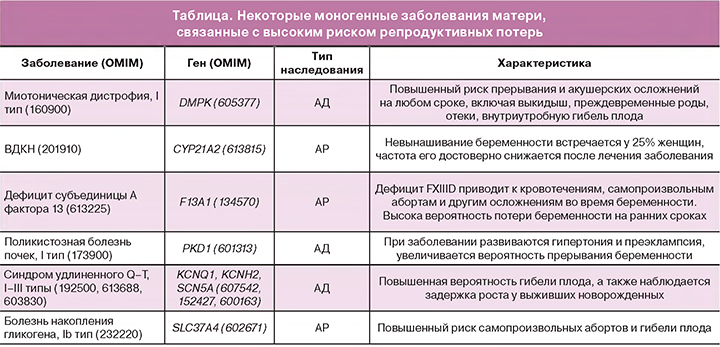
Повышенный риск самопроизвольных абортов и гибели плода
Важной проблемой также являются летальные фенотипы у плода, обусловленные моногенными заболеваниями: аутосомно-рецессивные заболевания (α-талассемия; синдром множественных птеригиумов, летальный тип; галактосиалидоз; мукополисахаридоз, VII тип), аутосомно-доминантные заболевания (танатофорная дисплазия; несовершенный остеогенез, II тип; ахондроплазия; туберозный склероз, I тип), Х-сцепленные заболевания (синдром недержания пигмента (Блоха–Сульцбергера); синдром Гольца (фокальная кожная гипоплазия); синдром Ретта; синдром иммунной дисрегуляции, полиэндокринопатии и энтеропатии).
Fridman et al. изучили 6447 последовательностей экзома здоровых, генетически неродственных европейцев из Голландии и Эстонии и подсчитали, что почти все люди (>85%) являются носителями по крайней мере одного патогенного или вероятного патогенного варианта со средним значением не менее 1,3 для тяжелого аутосомно-рецессивного заболевания (АР) и 2,2 для любого АР [17]. По оценкам, у кровнородственных пар, состоящих из двоюродных братьев и сестер, риск зачатия ребенка с АР-заболеванием в 16 раз выше, чем у неродственных пар, что означает 3400 новорожденных с тяжелым АР-заболеванием на 100000 рождений у двоюродных братьев и сестер (3,4%) [18]. Интересно отметить, что, как и ожидалось, риски для более отдаленных родственников постепенно снижались, а риск для троюродных братьев был таким же, как и для не состоящих в кровном родстве парах [17].
В контексте генетического тестирования для прогнозирования риска заболевания клиническая полезность особенно зависит от наличия вмешательств, которые могут эффективно снизить риск среди людей, отнесенных к группе высокого риска. Сегодня многие генетические тесты и оценки позволяют делать «прогноз без обещаний», то есть не существует известного способа снижения связанного с этим риска. Наконец, связанные с этим этические, правовые и социальные вопросы должны быть рассмотрены для всех тестов, выявляющих генетическую информацию, особенно в отношении сохранения генетической идентичности и сохраняющейся проблемы вариантной интерпретации (рис. 1).
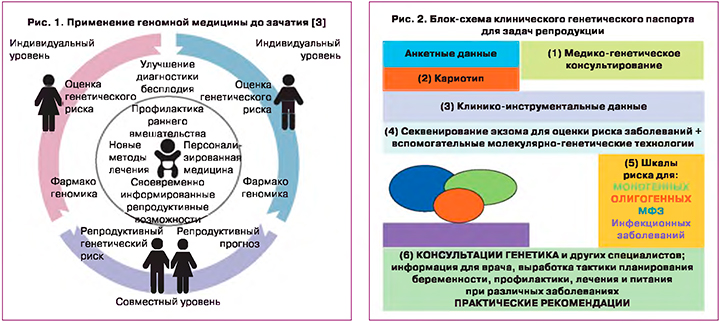
Геномная оценка до зачатия может обеспечить клиническую полезность как на индивидуальном, так и на супружеском уровнях. На индивидуальном уровне геномная информация может быть использована для выявления причин бесплодия, а также риска для здоровья при состояниях, не связанных с фертильностью. Конкретные варианты могут быть использованы для подбора фармакологического лечения либо бесплодия, либо общего состояния здоровья. На уровне супружеской пары объединенная геномная информация может быть использована для выявления потенциального риска передачи рецессивных генетических заболеваний потомству, что позволяет своевременно принимать обоснованные решения о репродуктивных стратегиях [3].
В нашем исследовании по возможности оказания медицинской помощи на примере семьи с наследственной патологией показано успешное использование различных молекулярно-генетических технологий [19]. Данный пример демонстрирует необходимость внедрения экзомного секвенирования и нового алгоритма преконцепционного обследования семей с использованием всего арсенала молекулярно-генетических методов, включая секвенирование нового поколения как метод первого звена при планировании беременности, а также метода пренатального генетического тестирования (ПГТ) и неинвазивного пренатального тестирования (НИПТ) для последующего мониторинга беременности. Мы предлагаем для предиктивного тестирования применять разработанный нами комплекс тестов, в основе которого экзомное секвенирование – клинический генетический паспорт. Тестирование включает следующие направления: сбор анкетных данных, медико-генетическое консультирование, кариотипирование, исследование на носительство моногенных заболеваний (экзом), с последующим планированием беременности, дифференциальной диагностики и лечения, подтверждающей диагностики (рис. 2).
Преконцепционный (до зачатия) генетический скрининг
Важнейшим вопросом медицины является решение проблем репродукции, а ключевым инструментом для этого является внедрение преконцепционного генетического скрининга. Преконцепционный (до зачатия) скрининг – это генетическое исследование на носительство распространенных наследственных болезней, которое проводят парам, планирующим рождение ребенка.
Преконцепционный скрининг позволяет оптимизировать алгоритм ведения будущей беременности: 1) выбор диагностических процедур; 2) рекомендации по медицинскому прерыванию; 3) консультирование; 4) междисциплинарный подход; 5) может быть использован на этапе планирования беременности: 6) донорство; 7) ПГТ; 8) снижает количество перинатальных потерь; 9) важен при психологической поддержке будущих родителей (позволяет уменьшить количество самообвинений среди тех, кто столкнулся с самопроизвольным прерыванием беременности, в том числе из-за моногенных заболеваний); 10) безопасен и прост для пациента.
Важно отметить, что преконцепционный скрининг с помощью NGS позволяет тестировать на носительство нескольких рецессивных заболеваний одновременно. Можно проводить тестирование, как пары, так и отдельных лиц независимо от особенностей родословной или географического происхождения. Особенно широко внедряется это тестирование в клиниках по лечению бесплодия (предлагают тестирование до зачатия). При этом, специалисты ЭКО все больше обращают внимание на необходимость этического руководства для подобного тестирования [20]. Хотя этические вопросы и не являются задачами данного исследования, следует отметить несколько ключевых моментов: явно ли возможные выгоды перевешивают возможный вред, недостатки? И если да, то, при каких заболеваний, и при каких изменениях в генах [20]? Все это очень важно для того, чтобы избежать передачи патогенных вариантов в генах, приводящих к болезням своим детям, используя технологию ПГТ-М. В некоторых случаях скрининг является обязательным и контролируется законами. Например, супружеские пары в Иране, Саудовской Аравии и на Кипре по закону обязаны участвовать в добрачном обследовании на наличие гемоглобинопатий [21]. Следует отметить, что в европейских странах 2,6% родов достигается с помощью вспомогательных репродуктивных технологий [22]. Пары, которые уже занимаются лечением бесплодия, в целом позитивно настроены в отношении скрининга на носительство, поскольку его можно легко добавить к запланированному циклу ЭКО и увеличить шансы на рождение здорового ребенка. Так, в 2015 г. Martin et al. опубликовали исследование, в котором 138 пар, обратившихся за вспомогательной репродукцией с использованием собственных гамет, участвовали в скрининге на более чем 600 заболеваний до зачатия. Семь пар были идентифицированы как пары-носители. Эти пары получили генетическую консультацию, и им было рекомендовано выбрать ПГТ-М [23]. Несмотря на это уникальное исследование, необходимы более масштабные последующие исследования, чтобы описать фактическое использование ПГТ-М после скрининга. Основные положения таких исследований отражены в директивах Европейского общества репродуктивной генетики человека [24].
Заключение
Необходимо внедрение экзомного секвенирования в соответствии с концепцией клинического генетического паспорта репродукции, особенно на этапе преконцепции, наряду с уже расширяющимся неонатальным скринингом, что позволит повысить рождаемость, обеспечить сохранность беременности и увеличить репродуктивный потенциал населения России.



